
Epigenetics is a rapidly growing field of cancer research. This page discusses some of the basics of epigenetics and the link between epigenetic changes and cancer.
- Introduction to Epigenetics
- Epigenetic Modifications
- Epigenetic Changes Outside of Genes
- Causes of Epimutation
- DNA Methylation Changes Seen in Cancer
- Histone Modification Changes Seen in Cancer
- Epigenetics and Cancer Metastasis
- Epigenetics and Cancer Prevention
- Epigenetics and Cancer Detection
- Epigenetic Proteins as Cancer Biomarkers
- Epigenetics as a Cancer Drug Target
Introduction to Epigenetics
The biological basis of cancer was unknown until researchers made the link between the disease and defective genes. They found that changes (mutations), in the DNA sequence of specific genes led to the uncontrolled cell reproduction (cell division) seen in cancer.1 This led to the discovery of two major groups of genes linked to cancer. The first to be found were the oncogenes, which cause cancer when their activity is increased, and later, tumor suppressor genes were discovered. Tumor suppressors normally block cancer, but can help drive cancer when they are altered or eliminated. More recently, it has been realized that cancer can also be the result of epimutations – small chemical changes that alter gene activity without changing DNA sequences.
Almost every cell in our body contains the same DNA sequences, yet it is immediately clear that all our cells do not look and behave alike. Heart cells look and act very differently than those in the lungs, even though both kinds of cells contain the same DNA. The reason for the variety of activities seen in different cells is explained by epigenetics. Epi- is the Greek prefix for “above”. Epigenetic changes provide a way for cells to control and regulate gene activity without changing the genes permanently. Instead, epigenetic control relies on small, reversible, changes to the DNA and proteins that make up chromosomes. To understand epigenetics, it is necessary to understand the nature of DNA. DNA is composed of four types of chemical building blocks (nucleotides). The nucleotides share some parts with each other, but they each have unique component, called a ‘base’ – the bases in DNA are guanine (G), cytosine (C), adenine (A), and thymine (T). DNA is a spiraled, ladder-like structure, with pairs of bases in the middle. In cells, DNA does not float around freely; it is tightly packed in chromosomes – structures in which the DNA is wrapped tightly around proteins called histones.2 DNA is packed in this way for several reasons. One is that DNA molecules are very, very long – if it were all uncoiled, the DNA in a single cell would be about 6 feet (2 meters) long!3 It would be almost impossible to fit in a cell without some good organization. Another reason is to strictly control gene activity so that genes are active only when needed.
So how does a cell, turn on, or activate a gene it needs? First, the portion of DNA containing the gene needs to be unwound and the cell does this by modifying the histones. Enzymes either add or remove small chemical markers, causing the histones to loosen their hold on the DNA. When the DNA is made available, different proteins are able to stick to the target gene and use the encoded information. This process is tightly controlled because unregulated gene activity in cells can cause many problems, including the development of cancer.
Epigenetic Modifications
As stated above, epigenetic control depends on small chemical changes to DNA or the proteins in chromosomes. There are several types of epigenetic modification. The major ones are described below.
1. DNA Modifications
The most common DNA modification is methylation. Methylation is the addition of a small chemical group - called a methyl group (-CH3) to specific bases, The bases are modified by enzymes called DNA methyltransferases (DNMTs). The most commonly altered base is cytosine. The addition of many methyl groups to a gene usually results in the gene being deactivated (silenced). There are several ways that the addition of these small groups can lead to the shutting off a gene. The first is that the methyl groups prevent enzymes that read DNA from recognizing and binding to the target gene, and the second is that the methyl groups on the DNA recruit DNA-binding proteins that block gene activation.1 Genes can not be 'read' if the right proteins can not stick to them. DNA methylation, like histone modification, is strictly controlled and problems arise when normal DNA methylation patterns are changed. Some tumor cells have been discovered to be globally under-methylated - they have decreased DNA methylation across many of their genes.4 Because methylation is linked to the activity of genes, changes in the methylation of single genes can also lead to cancer. Genes that drive cells to reproduce can be made overly active or genes that normally prevent cells from growing out of control can be turned off.
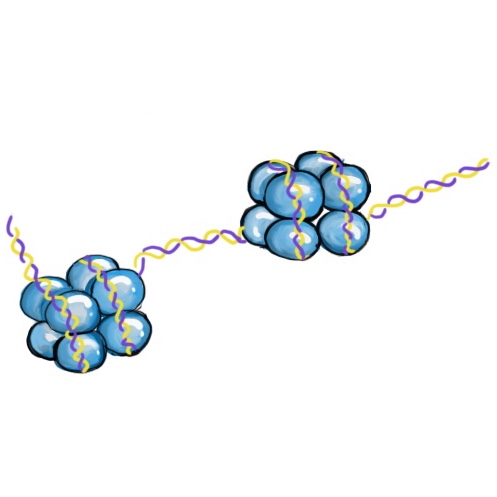
DNA wrapped around groups of histone proteins. Each group of 8 histones is called a nucleosome.
2. Histone Modifications
DNA is organized in structures called nucleosomes, which resemble beads on a string. Each nucleosome is composed of DNA wrapped around proteins called histones. There are 8 histone proteins in each nucleosome (2 copies each of histones H2A, H2B, H3, and H4).2 The histones are arranged in the nucleosomes so that the ends of the proteins (their “tails”) stick out from the center of the structure. The protein tails are a main site of histone modification. Like all proteins, histones are made up of amino acids. Amino acids located in the tail of the histones are targets for enzymes that attach or remove chemical markers. In particular, the amino acids lysine and serine are common targets of modification.
Compared to DNA methylation, histone modification is a relatively complicated process. DNA methylation involves only two types of enzymes, those that add methyl groups to cytosine and those that remove them. There are three different types of histone modification and each has multiple types of associated enzymes. The three types of histone modifications:
- Histone phosphorylation: The addition of phosphate groups to amino acids in the histone proteins.
- Histone methylation: The most complex of the three types. Methyl groups are added to amino acids in the histone proteins. More than one kind of amino acid can be modified, and individual amino acids can have more than one methyl group added. The complexity of these steps is reflected in the many enzymes associated with histone methylation. Each addition or removal process requires a specific enzyme.
- Histone acetylation: The addition or removal of an acetyl group to the amino acid lysine. Often found on the tails of histones. Histone acetylation causes the histone to become less positively charged, and the attraction between the histone and its bound DNA is weakened.5 This results in increased gene activity because the DNA is looser and more available.
The small changes described above alter the shape of nucleosomes and change the way DNA interacts with that particular “bead”. Histone modification is a very complex process and is tightly controlled. When histone modification patterns are altered, it can lead to unregulated activity or silencing of genes. Eventually, this can lead to the death of that cell, or more seriously, to cancer.
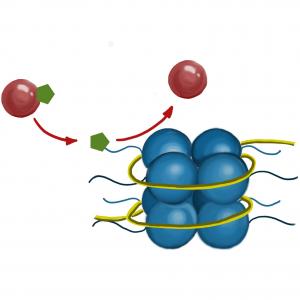
The diagram shows an enzyme (red ball) adding a chemical group (green hexagon) to the tail of a histone protein.
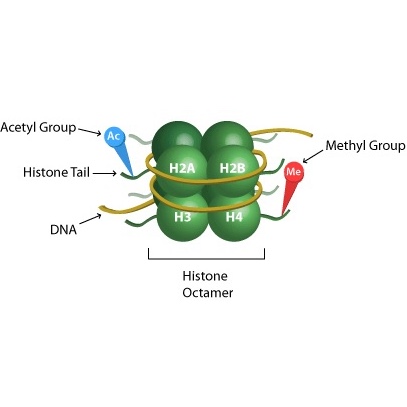
A more detailed view of nucleosome structure, showing targets of methylation and acetylation on the histone 'tails'
Epigenetic Changes Outside Genes
It is now understood that epigenetics plays a role in the development of cancer (carcinogenesis). As detailed above, abnormal epigenetic modifications in specific oncogenes and tumor suppressors genes can result in uncontrolled cell growth and division. However, abnormal epigenetic modifications in regions of DNA outside of genes can also lead to cancer. Protein-coding genes make up a small portion of our DNA. Humans have approximately 20,000 genes. The DNA occupied by our genes takes up less than 10% of all our DNA.6 The remaining 90% is made up of non-coding sequences. The non-coding sequences include regulatory sequences and regions of DNA that serve to provide structure to chromomes. Changes in these regions have been linked to several diseases, including cancer. It is important to note that epimutations that occur in a cell are passed on to the daughter cells formed when that cell reproduces by mitosis.
Causes of Epimutations
What causes abnormal epigenetic modifications? The environment and human behavior are main causes. Poor diet, lack of exercise, drugs, exposure to environmental chemicals or radiation – these all have the potential to cause epimutations which can lead to cancer. Smoking cigarettes, for example, has been shown to affect DNA methylation patterns across multiple organ systems. The affected genes have been linked to several major diseases including cancer, osteoporosis, chronic obstructive pulmonary disease (COPD), cardiovascular disease, and rheumatoid arthritis.7 In addition behavior and environment, the set of genes a person has definitely plays a role.
A very important fact about epigenetics is that the changes to genes can be passed down from parent to child. The term for this is “transgenerational inheritance”. In other words, some epigenetic effects are inherited. One of the first studies to show this transgenerational epigenetic inheritance was done in mice. In the study, a exposing a parent to a specific odor affected the behavior and sensory neurons in later generations.8 This study was groundbreaking because it showed that we can be affected by the experiences of our parents and grandparents. The exact mechanism behind this remains unclear.
Transgenerational inheritance was thought to be impossible due to something called germline reprogramming.9 This is a process in which the DNA of reprouctive cells (sperm and egg cells) is “reset” and all epigenetic marks and modifications are erased. In theory, this would eliminate the possibility of transfer of epigenetic events between generations. However, this process isn’t as complete as we once believed. Researchers identified the DNA methylation spots in germ cells and found that although most of the genome does get demethylated, a significant number of genes retain their epigenetic marks.9 Why this happens is up for debate, but some researchers think that keeping some parental epigenetic marks may increase the offspring’s chance of survival. A practical result of this new knowledge is that future parents need to consider the possible impact of their behaviors (diet, tobacco and alcohol use, etc.) on their unborn children.
Genetics is a major factor in determining someone’s risk of developing cancer. People with certain genetic mutations carry a relatively high risk of developing cancer during their lifetimes and these mutations can often be passed on to offspring. It is thought that about one-in-twenty to one-in-ten (5%-10%) of all cancers are due to inherited mutations.10 Now that we know epigenetic changes can also be inherited, those types of changes should be considered in future studies of cancer risk.
DNA Methylation Changes Seen in Cancer
Cancer cells often have a different epigenome, or epigenetic profile, than normal cells. DNA with less than normal amounts of DNA methylation are said to be hypomethylated. DNA with more methylation is said to be hypermethylated. A cancer cell’s epigenetic profile is typically characterized by decreased methylation across much of the genome (global DNA hypomethylation).11 The decreased methylation affects the activity of large numbers of genes. Because methylation is associated with decreased gene activity, the overall effect of hypomethylation is to increase the activity of the affected genes. If genes involved in cell growth have decreased methylation, the increased activity, and resulting cell division can lead to the development of cancer. As noted, DNA methylation changes do not have to be within protein-encoding genes to be important. Changes to DNA sequences that function as gene regulators can also cause problems.
Although the DNA of cancer cells is most commonly hypomethylated, the opposite can also be true.12 DNA hypermethylation in cancer cells tends to be limited to very specific regions (‘hot spots’). This sites affected vary by cancer type.The effect of increased DNA methylation is the opposite of hypomethylation. Hypermethylated genes tend to show decreased activity. DNA hypermethylation in cancer cells is frequently found at tumor suppressor genes; genes that function to repair DNA and control cell division. When tumor suppressor genes are silenced by increased methylation, the decrease in their activity can result in cancer development.
How do these small changes affect cancer growth? Cells with hypermethylated tumor suppressor genes are likely to grow faster than cells with normally methylated tumor suppressor genes. However, this doesn’t explain why specific sequences are found to be hypermethylated in some cancer cells but not others. The exact reason and mechanisms behind this is still unclear but, it may have something to do with how cell populations evolve over time. It is likely that the hypermethylation occurs to many genes in different cells. SOME of those changes will give those cells an advantage. Those cells will reproduce more rapidly, and take over the population.13 A good example is BRCA1, a gene linked to breast cancer, which is often found hypermethylated in breast and ovarian tumors but unmethylated in other types of tumors.14
Histone Modifications Seen in Cancer
Changes to the epigenetic modifications of histones also play an important role in the development of cancer. As previously mentioned,modifications of these proteins alter the interactions between the histones and the DNA. This changes the shape of the DNA-histone complexes (nucleosomes), and alters the way that other proteins can interact with the DNA.
The epigenome of cancer cells is typically marked by a loss of histone acetyl markers, due to increased histone deacetylation. The process of histone deacetylation is catalyzed by enzymes known as histone deacetylases or HDACs. As expected, increased activity of HDACs has been found in different types of cancer cells, making HDACs an important target of epigenetic cancer treatments.15
Histone methylation is also found to be affected in cancer cells. Histone methyltransferases (HMTs) are enzymes that carry out the addition of methyl groups to histones, while histone demethylases (HDMs) have the opposite function. In cancer cells, HMTs may be altered so that they are placing methyl groups at the wrong spot, often silencing tumor suppressor genes. HDMs can also be similarly affected, leading to increased activity of oncogenes. As previously mentioned, histone methylation is very complex. The effect of methylation on gene activity can differ depending on the specific amino acid affected. As such, methyl marks on histones are categorized as either activating or repressing, depending on their effect on gene activity. An additional complication is that some HDMs have been found to be able to remove both activating and repressing marks. This poses a challenge for people developing epigenetic therapies to target HDMs - their functions must be fully understood in order to know how the drugs will affect the cancer cells.
Epigenetics of Cancer Metastasis
The metastasis of cancer refers to the spread of the original (primary) tumor to a distant location in the body. Metastasis is a multi-step process: the cells must separate from the primary tumor, travel through the body to a new site through blood vessels or lymph vessels, reach a distant location, and then finally colonize the distant location to form a secondary tumor.16 Proteins have been identified that work to block cancer spread. These metastasis suppressors, which can inhibit any step of the process of metastasis. Metastatic cancer cells have been shown to epigenetically silence metastasis suppressors, often by hypermethylating these genes.17
The reasons why cancer cells metastasize is still not completely understood. Comparisons of the DNA sequences from metastatic cells and primary tumor cells were not always able to identify DNA sequence changes that could explain the difference between the cells. In 2017, researchers found an epigenetic basis for metastasis in at least one experimental model. This study examined pancreatic cancer cells from diseased patients and found that there were significant changes in the epigenome of metastatic cells, particular epigenetic changes that affected genes involved in cell migration.18
Epigenetics and Cancer Prevention
The main preventable causes of epimutations linked to cancer are environment exposures and behavior. Elimination or reduction of exposure to carcinogenic chemicals such as those found in tobacco products would likely reduce epimutations and related cancers. Other chemicals and drugs have also been found to cause epimutations, notably alcohol, which causes both DNA methylation and histone modification.19 A diet high in cruciferous vegetables, such as broccoli, cabbage, cauliflower, and kale, has been linked to a lower risk of developing prostate cancer.20 This is thought to be due to the action of several plant chemicals (phytochemicals), including sulforaphane and diindolylmethane (DIM). When researchers exposed prostate cancer cells to DIM, HDAC activity was reduce and increased amounts of the p21 tumor suppressor protein were produced.
Epigenetics and Cancer Detection
Current methods for detecting cancer use imaging techniques (X-rays, PET scans, ultrasound, etc.) or direct samples of suspicious areas to detect tumor growth. These methods are useful in detecting cancer; however, many of them rely on the detection of abnormal growths which are already present. Epigenetic detection methods are being designed to surpass the sensitivity and specificity of current tests.
Methylation-specific PCR (MSP) is an epigenetic test to identify abnormal methylation patterns in DNA. Because cancer cells are characterized by abnormal DNA methylation, it should be possible to identify cancer-specific patterns. This analysis relies on bisulfite sequencing, a technique that can distinguish between cytosines which have been methylated from those which have not been altered. MSP is useful because of its sensitivity – it can detect a single tumor cell among tens of thousands of cells.21 Another benefit is that MSP, unlike biopsies, is non-invasive. MSP can detect cancer DNA methylation patterns in plasma, stool, sputum, and urine samples.
Liquid biopsies are a cancer detection that relies on the analysis of blood samples. Liquid biopsies rely on the presence of tumor cells and/or tumor DNA in the bloodstream. These can be analyzed for both DNA mutations and epigenetic changes.22 Liquid biopsies are used in a rapidly expanding number of diseases, including cancer. There are some drawbacks to the use of liquid biopsies for the detection of epigenetic changes seen in cancer. Of upmost importance is our limited knowledge of the epimutations linked to cancer. Because different types of cancers are associated with different epimutations, it is difficult to design a general test to detect cancer. Many researchers are working to identify and categorize epigenetic patterns linked to cancer, but the process is slowed by the fact that each identified change must be confirmed in many patients.
Epigenetics-Related Proteins as Biomarkers for Cancer
A biomarker is an indicator of a condition that is not easily directly measured. Examples of biomarkers include the use of blood cholesterol levels and blood pressure as indicators of heart disease. Most current research into epigenetic cancer detection methods relies on identifying and detecting specific epigenetic changes. Ideally, it would be great to identify a set of changes (or a protein) that is present in most or all types of cancer. A 2017 study has identified a protein which may serve as just such a marker. UHRF1 is a protein encoded by the UHRF1 gene. It binds to specific DNA sequences where it recruits DNMT1, a DNA methyltransferase. It also functions in the coordination of different epigenetic enzymes, such as DNMTs and HDACs, to maintain normal DNA methylation patterns and histone modification patterns.23 UHR1 has been shown to be highly active in many types of cancer which makes it a potential universal cancer biomarker.24 UHRF1 has been identified as a protein that promotes the development of cancer. Increased levels of UHRF1 causes global DNA hypomethylation by destabilizing DNMTs. Increased UHRF1 activity have been linked to tumor growth, metastasis, and resistance to radiotherapy in different types of cancer, including lung, liver, breast, pancreatic, colorectal, prostate, and kidney (renal). High levels of UHRF1 have also been linked to lower survival rates, increased resistance to therapy, and increased rates of recurrence. Abnormal UHRF1 activity can be detected in the early stages of tumor development, making it a good candidate for early diagnosis of cancer. Targeting UHRF1 may also improve the effectiveness of radiotherapy.
Cancer Treatments Targeting Epigenetic Changes
Most current cancer treatments, including radiotherapy and chemotherapy, are cytotoxic treatments. Their goal is to kill cancer cells. However, these treatments are limited in their effectiveness as they often damage or kill normal cells, and drug resistance frequently develops in the cancer cells. Due to these limitations, other types of treatments are being researched, including immunotherapies, and epigenetic treatments.
Epigenetic treatments or “epi-drugs” refer to drugs that reverse abnormal epigenetic modifications in cancer cells. The main benefit of epi-drugs is that they are not cytotoxic, their goal is to “reprogram” cancer cells.25 Unlike genetic mutations, epimutations are reversible, which allows for the possibility to reverse the epimutations in cancer cells. The targets of epigenetic therapy are the enzymes involved in epigenetic modifications, including HDACs, DNMTs and HDMs. Although any type of epigenetic enzyme is a potential target for epigenetic therapy, research has focused on enzymes that are less complex and well-studied. The two main types of epi-drugs being investigated are HDAC inhibitors and DNMT inhibitors.
Histone Deacetylase (HDAC) Inhibitors
Histone acetylation is the process by which enzymes add acetyl groups to histones. The result is often an increase in gene activity. The reverse process, histone deacetylation, has the opposite effect, leading to gene silencing. In many types of cancer cells, high histone deacetylase activities have been observed. The increase is linked to the silencing of tumor suppressor genes and DNA repair genes. The aim of HDAC inhibitors is to decrease HDAC activity and indirectly increase the activity of tumor suppressors.
Many HDAC inhibitors are currently being researched and several of these have been approved for clinical use including vorinostat (Zolinza®), romidepsin (Istodax®), bellinostat (Beleodaq®), and panobinostat (Farydak®).26 These epi-drugs work by binding to HDACs, blocking their ability to remove acetyl groups from histones. These four drugs are pan-HDAC inhibitors - they target most or all HDACs. Although the approval of HDAC inhibitors is a step in the right direction, the epi-drugs have some limitations and disadvantages. One is that these drugs have only shown to be effective for treating blood (hematologic) cancers, including lymphoma, leukemia, and myeloma. Another drawback is that the drugs have side effects including fatigue and diarrhea. They are also toxic to bone marrow, and can reduce blood cell counts . The side effects are at least party due to their lack of specificity. By targeting all types of HDACs. They cause side effects by inhibiting HDACs (or other enzymes) necessary for normal cellular function. HDAC inhibitors designed to specifically target well-studied HDACs, could have better results and less side effects.
DNA Methyltransferase (DNMT) Inhibitors
As discussed above, DNA methylation is the process in which enzymes called DNA methyltransferases add methyl groups to bases in DNA. Because these changes affect the activity of the genes, DNA methylation plays a role in cancer development. Tumor suppressor genes are often heavily methylated and silenced. DNMT inhibitors have therefore been studied for their possible use as epi-drugs.27 In theory, DNMT inhibitors can reverse gene silencing and restore normal tumor suppressor function.
Two DNMT inhibitors have been shown to be effective as anticancer drugs and are currently approved for use in treating acute myeloid leukemia and myelodyplastic syndrome: azacytidine (Vidaza®) and decitabine (Dacogen®). When these drugs enter cells, they interact with DNA and inhibit any DNMTs that come along. The bound DNMT is unable to further methylate other regions of DNA and is ultimately destroyed. Although both epi-drugs are widely used in clinical treatment, they have several disadvantages. One drawback is their chemical instability; they are rapidly broken down, often in less than an hour and the drugs are changed into inactive compounds. Another downside is the toxicity of the drugs. They can cause DNA damage and lower immune function.
A new experimental DNMT inhibitor, zebularine, has been shown to be effective. Although it has not yet reached clinical trial stages, it is promising because it may improve upon the drawbacks of azacytidine and decitabine. Zebularine inhibits DNMTs and reverses gene silencing in cancer cells. It does so in a similar method to azacytidine and decitabine by interacting with DNA.28 When DNMTs encounter DNA with zebularine stuck to it, a a very stable complex is formed,’trapping’ the DNMT and blocking methylation. Zebularine offers several advantages over the two approved DNMT inhibitors. It is more stable in the body and it appears to be much less toxic.
Combination Epigenetic Drug Therapy
Complex diseases such as cancer often prove difficult to treat. Cancers also frequently develop resistance to any individual treatment. Combination therapy has emerged as a method to treat cancer more effectively. Combination therapy refers to the use of multiple treatments at the same time. For cancer patients, this often involves some combination of surgery, epigenetic drugs, radiotherapy, chemotherapy, and targeted therapies.
The combination of epigenetic drugs and radio/chemotherapy appears to be useful as each therapy may be able to compensate for the other’s disadvantages. Radiation and chemotherapy are effective in killing cancer cells and slowing growth; however, these treatments are often not be enough on their own. If all the cancer cells are not killed, then recurrence is possible. Combination treatments that include epigenetic drugs may increase sensitivity to chemotherapy and radiation therapy. 29
Combination epigenetic therapy is also a possibility. This involves the use of two or more different types of epigenetic drugs, including HDAC inhibitors and DNMT inhibitors. A significant benefit to combined epigenetic therapy is that it allows the use of lower drug doses. This could reduce side effects and increase effectiveness. Research with acute myeloid leukemia cells showed that combined decitabine-vorinostat treatment resulted in increased cancer cell death.30 Combining HDAC and DNMT inhibitors can result in increased histone acetylation compared to the use of HDAC inhibitors alone. The increase can be explained by the “communication” that occurs between DNA methylation and histone modification. The effect of DNA methylation is boosted by histone modification. Proteins that bind to methyl groups on DNA have the ability to recruit other enzymes, including HDACs, further decreasing gene activity.31
Limitations and Future Directions
Current epigenetic therapy is promising; however, several obstacles must be overcome to make it more effective. Although epigenetic drugs have shown to be effective against hematological cancer, a major limitation is their ineffectiveness against solid tumors. This is largely because of the environment of tumors typically have very low oxygen levels (low oxygen=hypoxia).32 Hypoxia occurs in tumors due to the presence of abnormal blood vessels, which cannot provide rapidly growing cancer cells with enough oxygen. The conditions alter the behavior of the tumor cells. Some arise which are characterized by aggressiveness, metastasis, and resistance to chemotherapy and radiation. Hypoxia results in the production of a different group of enzymes and proteins that take over control of gene activity. Additionally, tumor cells living in hypoxic conditions have an unusual epigenetic profile characterized by a decrease in active histone marks, an increase in repressive histone marks, and decreases in histone acetylation and DNA methylation. This is significant because current HDAC and DNMT inhibitors do not seem to be effective against tumor cells growing in hypoxic conditions. Research is required to better understand the epigenetic patterns of cancer cells living in hypoxic conditions. This information will help us to develop epigenetic drugs to target solid tumors.
Combination therapy can also be improved. This treatment approach relies on different types of cancer therapies working together; however, extensive research must be done to discover the best combinations of different drugs, proper dosing and timing. The development of new epigenetic drugs is also critical in improving cancer treatment. One of the main objectives is to design drugs with increased specificity. Ongoing research includes further exploration of epigenetic biomarkers linked to cancer and work to understand the epigenetic mechanisms of drug resistance in cancer cells. A major reason to study cancer epigenetics is inform the development of safe, effective treatments and to help prevent and detect cancer.
- 1ab Chen, Q. W., Zhu, X. Y., Li, Y. Y., & Meng, Z. Q. (2014). Epigenetic regulation and cancer. Oncology Reports, 31, 523-532. [PUBMED]
- 2ab Kouzarides, T. (2007). Chromatin Modifications and Their Function. Cell, 128, 693-705. [PUBMED]
- 3 Annunziato, A. (2008) DNA Packaging: Nucleosomes and Chromatin. Nature Education 1(1):26. [View]
- 4 Kulis, M., & Esteller, M. (2010). 2 – DNA Methylation and Cancer. Advances in Genetics, 70, 27-56. [PUBMED]
- 5 Bannister, A. J., & Kouzarides T. (2011). Regulation of chromatin by histone modifications. Cell, 21(3), 381-395. [PUBMED]
- 6 Powledge, T. M. (2014, August 12). How much of human DNA is doing something? Retrieved July 26, 2017 [View]
- 7 Joehanes, R., et al. (2016). Epigenetic Signatures of Cigarette Smoking. Circulation: Cardiovascular Genetics, 9(5), 436-447. [PUBMED]
- 8 Dias, B. G., & Ressler, K. J. (2013). Parental olfactory experience influences behavior and neural structure in subsequent generations. Nature Neuroscience, 17, 89-96. [PUBMED]
- 9ab Heard, E., & Martienssen, R. A. (2014). Transgenerational Epigenetic Inheritance: myths and mechanisms. Cell, 157(1), 95-109. [PUBMED]
- 10 Genetic Testing for Hereditary Cancer Syndromes. (2013, April 11). Retrieved July 14, 2017. [View]
- 11 Sharma, S., Kelly, T. K., Jones, & P. A. (2010). Epigenetics in cancer. Carcinogenesis, 31(1), 27-36. [PUBMED]
- 12 De Bustros, A., Nelkin, B. B., Silverman, A., Ehrlich, G., Poiesz, B., & Baylin, S. B. (1988). The short arm of chromosome 11 is a "hot spot" for hypermethylation in human neoplasia. Proc Natl Acad Sci USA, 85(15), 5693-5697. [PUBMED]
- 13 Esteller, M. (2002). CpG island hypermethylation and tumor suppressor genes: a booming present, a brighter future. Oncogene, 21(35), 5427-5440. [PUBMED]
- 14 Esteller, M., et al. (2000). Promoter hypermethylation and BRCA1 inactivation in sporadic breast and ovarian tumors. J Natl Cancer Inst, 92(7), 564-569. [PUBMED]
- 15 Halkidou,K., et al. (2004). Upregulation and nuclear recruitment of HDAC1 in hormone refractory prostate cancer. Prostate, 59, 177–189. [PUBMED]
- 16 Mandal, A. (2014, October 08). What is Metastasis? Retrieved July 28, 2017 [View]
- 17 Li, Q. & Chen, H. (2011). Epigenetic modifications of metastasis suppressor genes in colon cancer metastasis. Epigenetics, 6(7), 849-852. [PUBMED]
- 18 McDonald, O. G. et al. (2017). Epigenetic reprogramming during pancreatic cancer progression links anabolic glucose metabolism to distant metastasis. Nature Genetics, 49, 367-376. [PUBMED]
- 19 Mahnke, A. H., Miranda, R. C., & Homanics, G. E. (2017). Epigenetic mediators and consequences of excessive alcohol consumption. Alcohol, 60, 1-6. [PUBMED]
- 20 Beaver, L. M., et al. (2012). 3,3′-Diindolylmethane, but not indole-3-carbinol, inhibits histone deacetylase activity in prostate cancer cells. Toxicology and Applied Pharmacology, 263(3), 345-351. [PUBMED]
- 21 Zhu, J., & Yao, X. (2008). Use of DNA methylation for cancer detection: Promises and challenges. The International Journal of Biochemistry and Cell Biology, 41, 147-154. [PUBMED]
- 22 Alix-Panabières, C. & Pantel, K. (2013). Circulating Tumor Cells: Liquid Biopsy of Cancer. Clinical Chemistry, 59(1), 110-118. [PUBMED]
- 23 Unoki, M., Brunet, J., & Mousli, M. (2009). Drug discovery targeting epigenetic codes: the great potential of UHRF1, which links DNA methylation and histone modifications, as a drug target in cancers and toxoplasmosis. Biochem Pharmacol, 78, 1279-1288. [PUBMED]
- 24 Ashraf, W. et al. (2017). The epigenetic integrator UHRF1: on the road to become a universal biomarker for cancer. Oncotarget. [PUBMED]
- 25 Ronnekleiv-Kelly, S. M., Sharma, A., & Ahuja, N. (2017) Epigenetic therapy and chemosensitization in solid malignancy. Cancer Treatment Reviews, 55, 200-208. [PUBMED]
- 26 Ceccacci, E., & Minucci, S. (2016). Inhibition of histone deacetylases in cancer therapy: lessons from leukaemia. British Journal of Cancer, 114(6), 605-611. [PUBMED]
- 27 Gnyszka, A., Jastrzebski, Z., & Flis, S. (2013). DNA Methyltransferase Inhibitors and Their Emerging Role in Epigenetic Therapy of Cancer. Anticancer Research, 33(8), 2989-2996. [PUBMED]
- 28 Champion, C. et al. (2010). Mechanistic Insights on the Inhibition of C5 DNA Methyltransferases by Zebularine. PLoS One, 5(8), e12388. [PUBMED]
- 29 Pajonk, F., Vlashi, E., & McBride, W. H. (2010). Radiation Resistance of Cancer Stem Cells: The 4 R’s of Radiobiology Revisited. Stem Cells, 28(4), 639–648. [PUBMED]
- 30 Young, C. S., Clarke, K. M., Kettyle, L. M., Thompson, A., & Mills, K. I. (2017). Decitabine-Vorinostat combination treatment in acute myeloid leukemia activates pathways with potential for novel triple therapy. Oncotarget. [PUBMED]
- 31 Jones, P. L., et al. (1998). Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nature Genetics, 19, 187-191. [PUBMED]
- 32 Ramachandran, S., Ient, J., Göttgens, E., Krieg, A. J., & Hammond, E. M. (2015) Epigenetic Therapy for Solid Tumors: Highlighting the Impact of Tumor Hypoxia. Genes, 6, 935-956. [PUBMED]
